Why Choose the K. lactis Protein Expression Kit?
The K. lactis Protein Expression Kit from NEB enables high yield protein expression in yeast.
Many types of protein can be secreted from yeast cells. As a general rule, those that tend to secrete best are proteins that are also secreted by their native host (e.g. glycosidases, serum albumins, cytokines, etc). However, there are numerous examples in the literature of normally non-secreted proteins that have been successfully secreted from various yeasts. When in doubt, it is always best to try secreted expression. Intracellular protein expression in yeast is also possible for a wide range of proteins and is a great alternative to bacterial protein expression.
K. lactis strain GG799 (as supplied in the NEB Kit) is a haploid (α) wild-type industrial isolate that has no genetic markers. It was originally chosen as a host strain in the food industry because of its ability to grow to very high cell density and to efficiently secrete heterologous proteins.
Advantages:
- High yield protein expression
- Multiple protein expression
- Rapid high cell density growth
- Methanol-free growth media
- No background gene expression during E. coli cloning steps: Clone and express genes toxic to E. coli
- Easy and fast cell transformation: Competent K. lactis cells included
- No expensive antibiotics required
- Easy-to-use protocols for those inexperienced with yeast
- Attractive commercial sublicensing (no license required for research use, even in a commercial setting)
Avoid toxicity problems in E. coli
The K. lactis Protein Expression Kit vector (pKLAC2) contains a variant of the strong K. lactis LAC4 promoter (PLAC4-PBI) for expression of a desired gene in K. lactis. The major advantage of the PLAC4-PBI promoter is that it is transcriptionally silent while in E. coli. In contrast, the wild-type PLAC4 promoter shows background transcriptional activity in E. coli which can be detrimental to the process of assembling or amplifying expression constructs in E. coli prior to their introduction into yeast cells. This is especially problematic if the cloned gene of interest encodes a translated product that is toxic to E. coli cells. Therefore, pKLAC2 is well-suited for the cloning and yeast expression of genes encoding proteins that are toxic or otherwise detrimental to bacteria.
High yield, lower cost protein expression
Integrative expression with no auxotrophic markers: pKLAC2 is an integrative expression vector that inserts into the promoter region of the LAC4 locus of the K. lactis genome upon its introduction into K. lactis cells. While K. lactis episomal plasmids do exist, they can present some problems for large-scale protein production. For example, plasmids are easily lost by cells in the absence of a selection. For large-scale fermentation, plasmid selection using antibiotics can be too costly, and selection using an auxotrophic marker can reduce yields. While auxotrophic markers have historically been used for genetic manipulation of yeasts, they are not always desirable to achieve maximum protein expression. In some cases, an auxotrophy (e.g. uracil) can cause a significant reduction in the strain’s ability to produce a heterologous protein even if exogenous uracil or uridine is provided in the growth medium. Integrative expression vectors are attractive because they insert into the genome, thus becoming part of the host chromosome, and are therefore quite stable even in the absence of selection.
Acetamide selection: There are two main advantages to acetamide selection: cost and selection of multiple integrants. Acetamide is significantly less expensive than antibiotics. Additionally, acetamide selection enriches transformant populations for cells that have integrated multiple tandem copies of the pKLAC2 expression vector. Multi-copy integrants are desirable because they often produce more recombinant protein than single integrants. Acetamide acts as a nitrogen source. A cell transformation mixture (containing a population of cells that are either transformed or untransformed by vector pKLAC2) is spread onto yeast carbon base (YCB) medium agar containing 5 mM acetamide. YCB medium contains all of the nutrients and carbon source required for K. lactis cells to grow, but lacks a nitrogen source. The acetamide provided in the medium can be utilized as a source of nitrogen only if it is broken down to ammonia by the enzyme acetamidase (expressed from the amdS gene present on pKLAC2). Therefore, only transformed cells are able to grow into colonies.
Flexible cloning strategies
For cytosolic protein expression: The gene of interest can be cloned into pKLAC2 in a manner that places it downstream of PLAC4-PBI without the mating factor secretion domain being present. Genes expressed in this manner must include a methionine as their first codon, to initiate translation.
For co-expression of multiple proteins: An easy method for construction of K. lactis strains that co-express multiple proteins has been described by Read et al (3). This method exploits the tendency of pKLAC-series vectors to tandemly insert into the LAC4 promoter locus at exceptionally high frequency. This phenomenon permits separate pKLAC-series vectors, each containing a different heterologous gene, to co-integrate at the LAC4 locus generating cells that co-express each gene. This method has been used to create K. lactis strains that produce up to four different heterologous proteins after only one cell transformation reaction (3).
For tagged proteins: A PCR method for the addition of a carboxy-terminal HA tag to a secreted protein is described in the kit manual. This example can be adapted for the addition of other antibody epitope tags (e.g. FLAG, c-myc or His tag). Additionally, a chitin-binding domain has been used as a tag for capture of secreted proteins onto inexpensive chitin beads directly in spent medium (1).
For expression of proteins for structural analysis: The K. lactis Protein Expression Kit has been used to express 13C and 15N labeled proteins for structural analysis using NMR spectroscopy (4). This system offers an efficient method of isotopic labeling at relatively low cost.
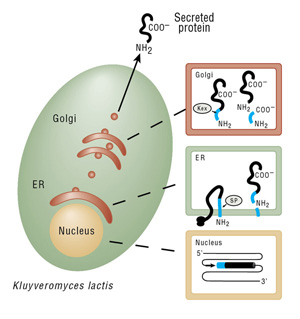
For an amino-terminal antibody epitope tag: A new forward PCR primer is used that contains an XhoI restriction site, the Kex protease cleavage site, the desired tag’s sequence and DNA homologous to the 5´ end of the desired gene or cDNA. After Kex protease processing of the expressed protein in the Golgi, a protein bearing the desired tag at its amino-terminus is produced and secreted.
Ordering Information
| Product | NEB # |
| K. lactis Protein Expression Kit | E1000S |
References
- Colussi, P.A., et al. (2005). Characterization of a nucleus-encoded chitinase from the yeast Kluyveromyces lactis. Appl Environ. Microbiol. 71, 2862–2869
- Van Ooyen, A.J. et al. (2006). Heterologous protein production in the yeast Kluyveromyces lactis. FEMS Yeast Res. 6, 381–92.
- Read, J.D., et al. (2007). Acetamide selection of Kluyveromyces lactis cells transformed with an inegrative vector leads to high-frequency formation of multicopy strains. Appl. Environ. Microbiol. 73, 5088-5096.
- Sugiki, T., Shimada, I. and Takahashi, H. (2008). Stable isotope labeling of protein by Kluyvecromyces lactis for NMR study. J. BioMol. NMR, DOI 10.1007/510858-008-9276-9.
Notice to Buyer/User: The K. lactis Protein Expression Kit was developed from basic research at New England Biolabs, Inc. and DSM Biologics Company B.V. The buyer and user has a non-exclusive sublicense to use this system or any component thereof, including the K. lactis GG799 Competent Cells, for RESEARCH PURPOSES ONLY. A license to use this system for manufacture of clinical grade material or commercial purposes is available from New England Biolabs, Inc., or DSM Biologics Company B.V.

