
Using PaqCI® for Golden Gate Assembly - What Makes it a Special Addition to the NEB Assembly Portfolio?
Posted on Wednesday, April 7, 2021
By
Topic: Tips for the lab
But first, some background on Golden Gate Assembly
Before we get into the specifics about PaqCI, let’s delve into a bit of enzyme background to set the stage: Golden Gate Assembly is directly dependent on Type IIS restriction enzymes that have asymmetric DNA recognition sites and cleave outside of their recognition, or binding, sequences. We currently offer 50 Type IIS restriction enzymes, of which a subset have the necessary favorable characteristics for Golden Gate Assembly. Enzymes such as BsaI-HF®v2, BsmBI-v2, and BbsI-HF® have been our Golden Gate workhorses as they have historically been featured in published assembly protocols in the scientific literature, plus we have extensive experience using them here at NEB. During this time, and with input from our customers, we recognized how useful it would be to the research community to offer an enzyme with a 7-base recognition site for assembly, but one done the “NEB Way” – with fully optimized protocols and enzyme recommendations, for assemblies ranging from simple to complex, and at a reasonable price.
And that is where the PaqCI Golden Gate story begins, as we set down the path of finding a Type IIS enzyme with a highly desirable 7-base recognition site (see Figure 1), whose sites are less likely to be present in DNA sequences being assembled, yet is capable of the full range of assembly complexity that scientists require for their experiments. Through a collaboration between laboratories in our Research, Applications Development, and Production Departments, PaqCI was identified and cloned, and its expression was optimized. We also optimized a DNA activator for the enzyme, and developed protocols for single inserts, and simple-to-complex assemblies. All this – from discovery, to optimization, to being available for sale – despite the challenges we have all been facing conducting our science during a pandemic!
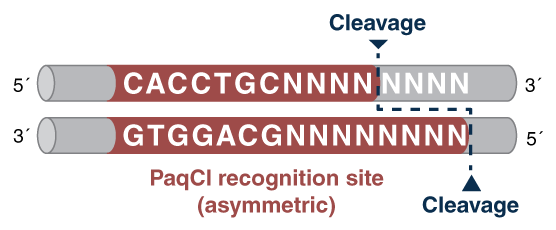
Figure 1: Type IIS enzymes recognize asymmetric DNA sequences and cleave outside of their recognition sequence
What is domestication? And why is PaqCI important when considering domestication?
Domestication refers to converting any DNA fragment that will be part of an assembly into “Golden Gate-ready” form - flanking the DNA at both ends with the Type IIS restriction sites that will direct the assembly and removing any internal sites for that enzyme that might be present in the DNA and are not tolerated well in GGA. For PaqCI it’s all about the 7- base recognition site; statistically any 7-base sequence will appear in any given DNA sequence less often than the 6-base sequence of the more commonly used Type IIS restriction enzymes. The Achilles’ Heel of Golden Gate Assembly has always been that internal sites significantly decrease assembly efficiency, as they would allow the finished construct to be susceptible to digestion by the restriction enzyme present in the assembly reaction at best, and could lead to incorrect and unwanted assemblies at worst.
This is less an issue when using Golden Gate for single insert cloning because the overall efficiency for single inserts is SO high; you would get your desired construct even if many of your successfully cloned inserts became linearized and did not transform well; simply by screening a few colonies from your transformation plate, you would find cloned single inserts. But typically, researchers are using Golden Gate for multiple inserts. And the more inserts you have—the greater the assembly complexity—the more you need maximal efficiencies. So, having an internal site for your chosen restriction enzyme is a big problem for Golden Gate Assembly.
There are proven methodologies for eliminating internal sites while domesticating DNA sequences. We have a video (below) that shows the options for domestication: (1) site-directed mutagenesis to eliminate an internal site in advance of the assembly reaction, or (2) designing an assembly junction point right at the internal restriction site with a base change to eliminate the site upon assembly.
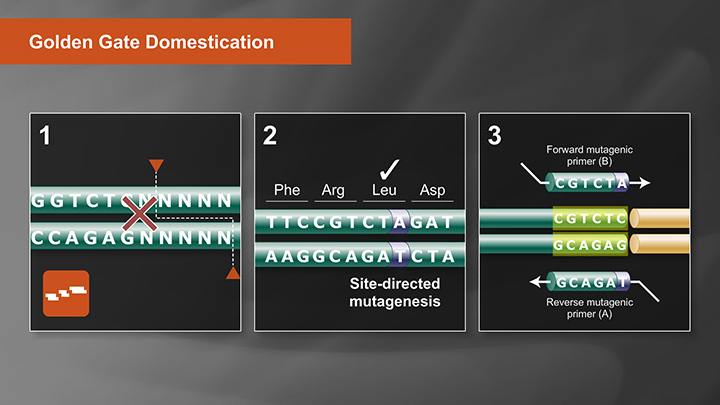
This video explains Domestication, or the removal of Type IIS cut sites naturally occurring in vector or insert sequences, as it relates to Golden Gate Assembly.
But all this sequence manipulation to deal with internal sites takes thought and time. Hence the attractiveness of a 7-base recognition site enzyme, which significantly decreases the probability of internal sites. And that is where PaqCI comes into the picture – a 7-base recognition restriction enzyme that has been optimized for Golden Gate assembly, and is supplied at a concentration that enables use for complex assemblies up to 20+ fragments. We’re very excited about this!
The mechanism of multi-site enzymes and why they benefit from the addition of an activator
Some enzymes have more intricate ways of interacting with their recognition sites in DNA than others. Most homodimeric enzymes, like the standard Type IIP restriction enzymes EcoRI and HindIII, have two identical subunits that bind cooperatively at the symmetric site with each subunit cutting one strand to result in a double-stranded cut. In contrast, multi-site enzymes like PaqCI have a more complex structure and mechanism. It is presumed that PaqCI utilizes multiple subunits to interact with two recognition sites in order to cleave a single target site. To make sure that PaqCI cuts all the sites during Golden Gate assembly, we supply an inert short oligonucleotide activator containing an extra PaqCI binding site, which functions in trans as an activator for PaqCI cleavage (see Figure 2).
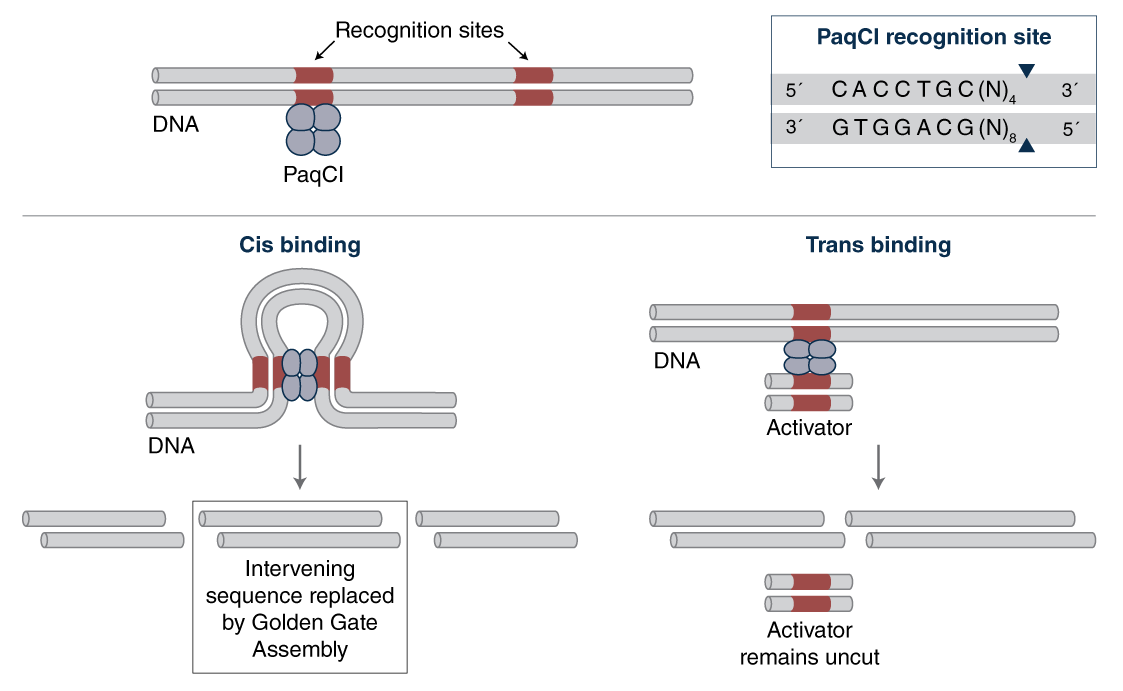
Figure 2: Presumed mechanism for how the PaqCI activator assures complete cutting via trans binding if needed
One would think that for Golden Gate, where by definition, every insert and every destination plasmid has its assembly active DNA fragment flanked by two sites, there would be no need for any added sites. But Golden Gate is a very dynamic process, with cutting and ligation taking place in the assembly reaction, and situations can arise where PaqCI binds and cuts sites on different DNA molecules, which means on each of those DNA molecules there would be another site remaining to be cut. So having an optimized number of extra sites available in the form of the PaqCI activator ensures that complete cutting in your assembly reaction occurs. It should be noted that the activator does not get cut or interact in any way with the assembly – it only provides a second binding site that can activate cutting. Different levels of complexity call for different levels of PaqCI and T4 DNA Ligase; in addition we have carefully optimized how much PaqCI and its activator are needed for different assembly complexities.
Now, it gets interesting when we think about what exactly constitutes the right amount activator. The cutting of DNA in a typical DNA restriction digest, where cut DNA remains cut, is different from what happens in Golden Gate assembly reactions, where sometimes cut sites are nonproductively reannealed and ligated, so that any one DNA cut site can require being cut more than once throughout the assembly reaction. This is why the optimal amount of the activator can be different from what is recommended for a standard restriction digest with PaqCI, where using 1 µl of the enzyme (10 U) requires 1 µl of the activator (20 pmoles). Because of the dynamic nature of GGA, these regenerated sites translate to less supplementary sites in the form of the activator being needed.
From over a thousand test assembly reactions, we can recommend just the right amount of PaqCI, activator, and T4 DNA Ligase for everything from simple single insert cloning to a complex 24-fragment assembly (see Table 1).
| Assembly Complexity | PaqCI | T4 DNA Ligase | PaqCI Activator3 |
|---|---|---|---|
| Single Insert Cloning (10 min 37°C) or Library Prep (60 min 37°C) | 5 U | 200 U | + 5 pmoles (1/4 µl 20 µM stock) |
| Simple to Moderate assembly1 (2-10 fragments) | 5-10 U | 200-400 U | +5 pmoles (1/4 ul 20 µM stock) |
| Complex assembly2 (11-20+ fragments) | 10-20 U | 400-800 U | +5-10 pmoles (1/4-1/2 µl 20 µM stock) |
- Based on 5 fragment assembly test system
- Based on 24 fragment assembly test system
- The activator solution is in a Mg-free buffer for best long-term storage. For short-term working stocks, if desired, dilute an appropriate amount in 1X T4 DNA ligase buffer to achieve more easily pipettable volumes (e.g., a four-fold dilution = 5 µM, 5 pmoles/µl activator.)
Table 1: Recommendations for PaqCI Golden Gate Assembly
As assembly reactions increase in complexity, more units of enzyme are required for maximal performance; our range is from 5 to 20 U of PaqCI paired with 200-800 U of T4 DNA Ligase. We also have recommendations for how much of our activator to add to each assembly reaction, within a range of 5-10 pmoles. We provide a 20 µM stock of the activator with each PaqCI enzyme, more than enough for all your assembly needs.
One note about the buffer requirements: CutSmart buffer is the best buffer to use for setting up a simple DNA digest with PaqCI, but for Golden Gate Assembly, there are better efficiencies achieved maximizing the PaqCI and T4 DNA ligase enzyme activities using T4 DNA Ligase Reaction Buffer.
New England Biolabs Golden Gate Assembly tools are here to help
Webtools make our scientific lives simpler by facilitating workflows, and our Golden Gate Assembly tool keeps to that tradition.
After designating the DNA fragments you would like to use for any given assembly, the tool can design optimal unique four base overhangs between the inserts that have been independently verified through T4 DNA Ligase fidelity studies to work at high fidelity. It will also automatically check your inserts for the presence of any internal sites that might affect the choice of Type IIS restriction enzyme to direct an assembly, or alert the user to remove such internal sites via domestication. The program will also automatically generate a set of primers for your inserts to add the flanking bases and recognition sites required either for amplicon generation of inserts to be directly used or for pre-cloning purposes. Finally, a report can be generated describing your full assembly with a color-coded graphical read out, your final assembly sequence, and descriptions of each junction between inserts.
In addition, there are also useful programs available under the “Utility” tab within the tool. Those programs can take an uploaded sequence and make suggestions for different desired insert or module design and can also provide you with vetted lists of what overhangs have been found to support high efficiencies and fidelities during Golden Gate Assembly. Together the Golden Gate Assembly tool makes assembly design easy, even for the first time user!
Check out our video outlining the Golden Gate Assembly workflow, our usage guidelines for GGA using PaqCI, or our protocol for GGA using PaqCI and T4 DNA Ligase.
NEB will not rent, sell or otherwise transfer your data to a third party for monetary consideration. See our Privacy Policy for details. View our Community Guidelines.
Products and content are covered by one or more patents, trademarks and/or copyrights owned or controlled by New England Biolabs, Inc (NEB). The use of trademark symbols does not necessarily indicate that the name is trademarked in the country where it is being read; it indicates where the content was originally developed. See www.neb.com/trademarks. The use of these products may require you to obtain additional third-party intellectual property rights for certain applications. For more information, please email busdev@neb.com.
Don’t miss out on our latest NEBinspired blog releases!
- Sign up to receive our e-newsletter
- Download your favorite feed reader and subscribe to our RSS feed
Be a part of NEBinspired! Submit your idea to have it featured in our blog.


